pyku.analyse#
Analysis module
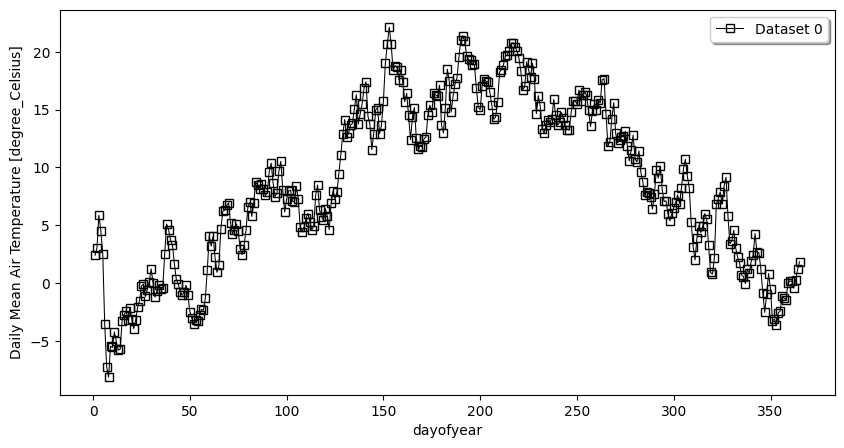
- pyku.analyse.daily_mean(*dats, var=None, ax=None, **kwargs)[source]#
Daily mean
- Parameters:
dats (
xarray.Dataset) – The input dataset(s)var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas').compute() ...: ds.ana.daily_mean(var='tas') ...: CPU times: user 2.15 s, sys: 422 ms, total: 2.57 s Wall time: 2.07 s
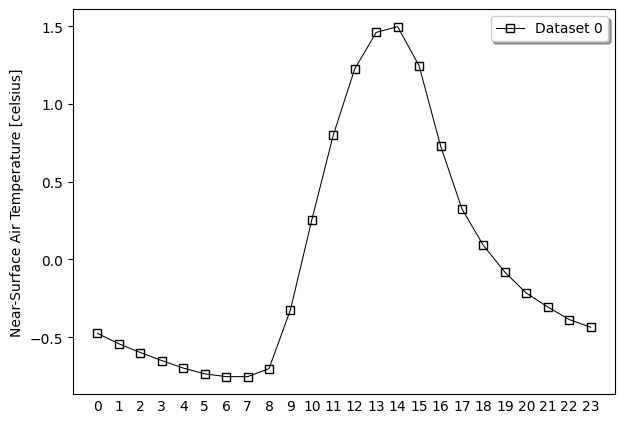
- pyku.analyse.diurnal_cycle(*dats, var=None, ax=None, **kwargs)[source]#
Diurnal cycle
- Parameters:
dats ([
xarray.Dataset, List[xarray.Dataset]]) – The input dataset(s)var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hourly-tas').compute() ...: ds.ana.diurnal_cycle(var='tas') ...: CPU times: user 8.31 s, sys: 9.73 s, total: 18 s Wall time: 16.5 s
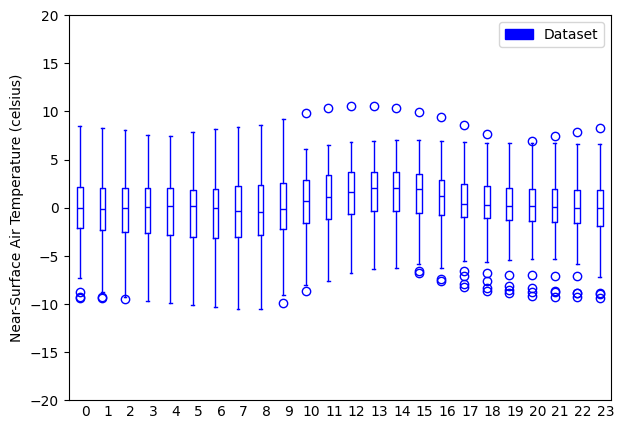
- pyku.analyse.diurnal_cycle_variability(dat1, dat2=None, var=None, ax=None, **kwargs)[source]#
Diurnal cycle variability
- Parameters:
dat1 (
xarray.Dataset) – The first dataset.dat2 (
xarray.Dataset) – Optional. The second dataset.var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()or func:matplotlib.pyplot.axes. In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hourly-tas') ...: ds.ana.diurnal_cycle_variability(var='tas', ylim=(-20, 20)) ...: CPU times: user 5.79 s, sys: 1.64 s, total: 7.44 s Wall time: 5.47 s
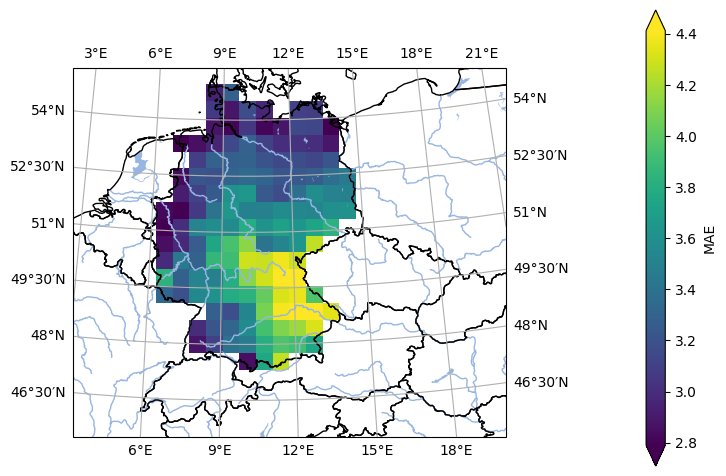
- pyku.analyse.mae_map(*dats, ref=None, var=None, crs=None, **kwargs)[source]#
Map of the mean absolute error.
- Parameters:
dats (
xarray.Dataset) – The input dataset(s).ref (
xarray.Dataset) – The reference dataset.var (str) – The variable name.
crs (
cartopy.crs.CRS) – The coordinate reference system.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Notes
If an argument is passed which is not recognized, no error message is thrown and the argument is ignored.
Example
In [1]: %%time ...: import pyku ...: ...: # Open reference dataset and preprocess as needed ...: # ----------------------------------------------- ...: ...: ref = pyku.resources.get_test_data('hyras')\ ...: .pyku.to_cmor_units()\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .sel(time='1981')\ ...: .compute() ...: ...: # Open model dataset and preprocess as needed ...: # ------------------------------------------- ...: ...: ds = pyku.resources.get_test_data('model_data')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .sel(time='1981')\ ...: .compute() ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.mae_map(ref=ref, var='tas', crs='HYR-LAEA-5') ...: CPU times: user 3.34 s, sys: 1.57 s, total: 4.91 s Wall time: 7.33 s
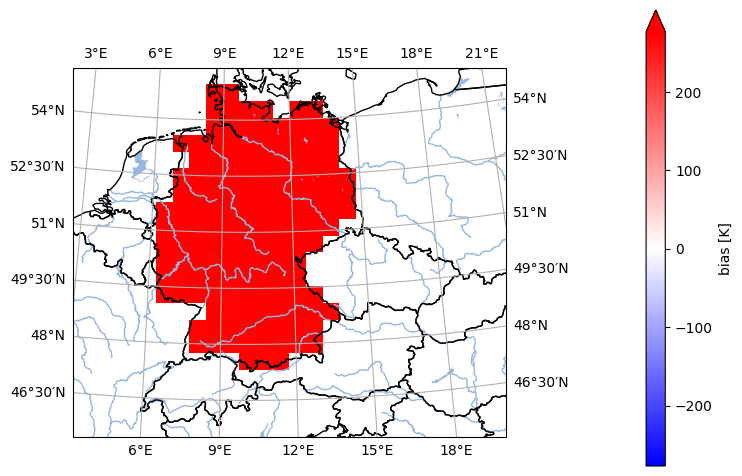
- pyku.analyse.mean_bias_map(*dats, ref, var=None, crs=None, **kwargs)[source]#
Map of the mean bias
- Parameters:
dats (
xarray.Dataset, List[xarray.Dataset]) – The input model dataset(s)ref (
xarray.Dataset) – The Reference/Observation datavar (str) – The variable name
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – Optional. The plot coordinate reference system.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Notes
If an argument is passed which is not recognized, no error message is thrown and the argument is ignored.
Example
In [1]: %%time ...: import pyku ...: ...: # Open reference dataset and preprocess as needed ...: # ----------------------------------------------- ...: ...: ref = pyku.resources.get_test_data('hyras')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .sel(time='1981')\ ...: .compute() ...: ...: # Open model dataset and preprocess as needed ...: # ------------------------------------------- ...: ...: ds = pyku.resources.get_test_data('model_data')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .sel(time='1981')\ ...: .compute() ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.mean_bias_map(dats=ds, ref=ref, var='tas', crs='HYR-LAEA-5') ...: CPU times: user 968 ms, sys: 293 ms, total: 1.26 s Wall time: 930 ms
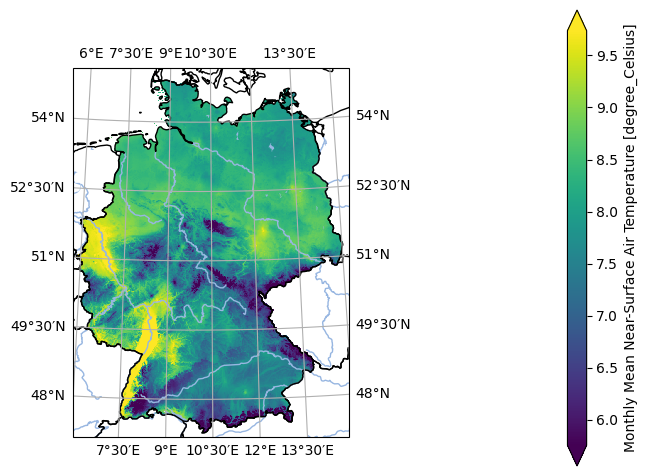
- pyku.analyse.mean_map(*dats, var=None, crs=None, same_datetimes=True, **kwargs)[source]#
Map of the mean
- Parameters:
dats (
xarray.Dataset, List[xarray.Dataset]) – The input dataset(s)var (str) – The variable name.
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – (Optional) Coordinate reference system of the plot.same_datetimes (bool) – (Optional) Defaults to
True. Select common datetimes between datasets.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Example
In [1]: %%time ...: import pyku ...: ds = ( ...: pyku.resources.get_test_data('hyras-tas-monthly') ...: .compute() ...: ) ...: ds.ana.mean_map(var='tas', crs='HYR-LAEA-5') ...: CPU times: user 653 ms, sys: 243 ms, total: 896 ms Wall time: 744 ms
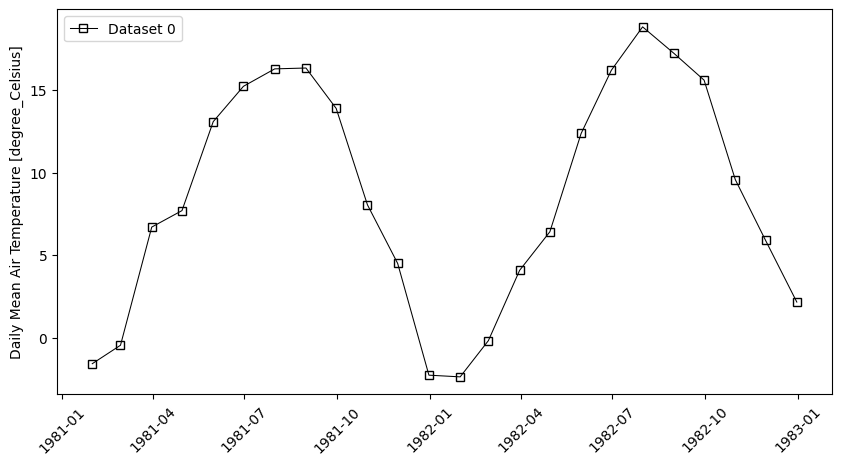
- pyku.analyse.mean_vs_time(*dats, var=None, time_resolution='1YS', ax=None, **kwargs)[source]#
Mean vs time
- Parameters:
*dats (
xarray.Dataset, List[xarray.Dataset]) – The input dataset(s).time_resolution (freqstr) – Time resolution (e.g. ‘1M’ or ‘6D’)
var (str) – The variable name.
ax (
matplotlib.pyplot.axes) – Optional. Matplotlib pyplot axis.**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hyras') ...: ds.ana.mean_vs_time(var='tas', time_resolution='1M') ...: CPU times: user 2.82 s, sys: 375 ms, total: 3.19 s Wall time: 3.21 s
- pyku.analyse.metadata(ds_mod, ds_obs)[source]#
Report on metadata
- Parameters:
ds_mod (
xarray.Dataset) – Model dataset.ds_obs (
xarray.Dataset) – Reference dataset.
- Returns:
reStructuredText report
- Return type:
str
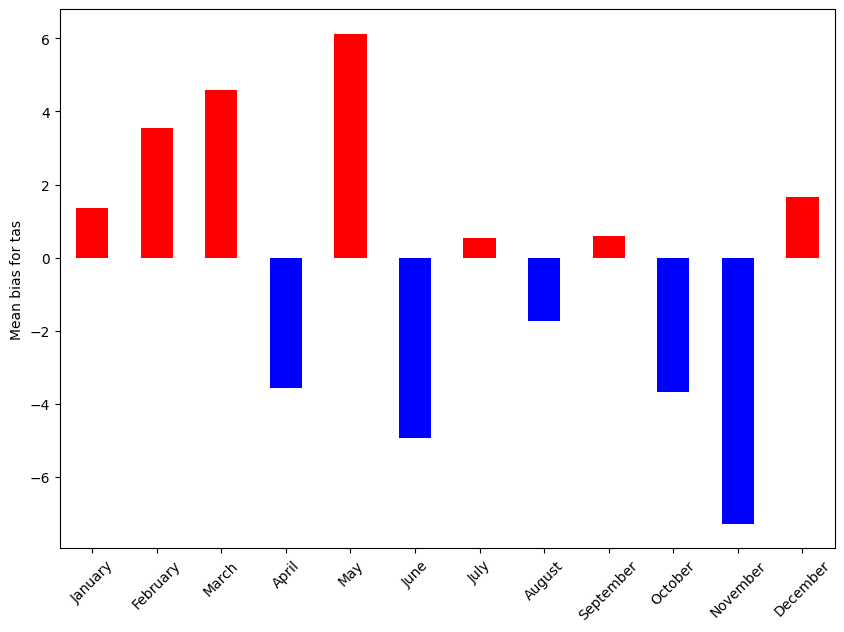
- pyku.analyse.monthly_bias(dat_mod, dat_obs, var=None, ax=None, **kwargs)[source]#
Monthly bias
- Parameters:
dat_mod (
xarray.Dataset) – The input dataset.dat_obs (
xarray.Dataset) – The reference dataset.var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: import pyku ...: ...: # Open reference data and preprocess as needed ...: # -------------------------------------------- ...: ...: ref = pyku.resources.get_test_data('hyras')\ ...: .pyku.to_cmor_units()\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .sel(time='1981')\ ...: .compute() ...: ...: # Open model data and preprocess as needed ...: # ---------------------------------------- ...: ...: ds = pyku.resources.get_test_data('model_data')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .sel(time='1981')\ ...: .compute() ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.monthly_bias(ref, var='tas') ...:
- pyku.analyse.monthly_bias_var(ds_mod, ds_obs, var, **kwargs)[source]#
Monthly bias variation
- Parameters:
ds_mod (
xarray.Dataset) – The input dataset.ds_obs (
xarray.Dataset) – The reference dataset.var (str) – The variable name.
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition) – (Optional) Coordinate reference system of the plot.**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ...: # Open, reproject ...: # --------------- ...: ...: ref = pyku.resources.get_test_data('hyras')\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .pyku.to_cmor_units()\ ...: .pyku.project('HYR-LAEA-50')\ ...: .sel(time='1981') ...: ...: # Open model, reproject, reset time labels, select the year ...: # --------------------------------------------------------- ...: ...: ds = pyku.resources.get_test_data('model_data')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .sel(time='1981') ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.monthly_bias_var(ref, var='tas') ...: CPU times: user 1.55 s, sys: 333 ms, total: 1.88 s Wall time: 1.21 s
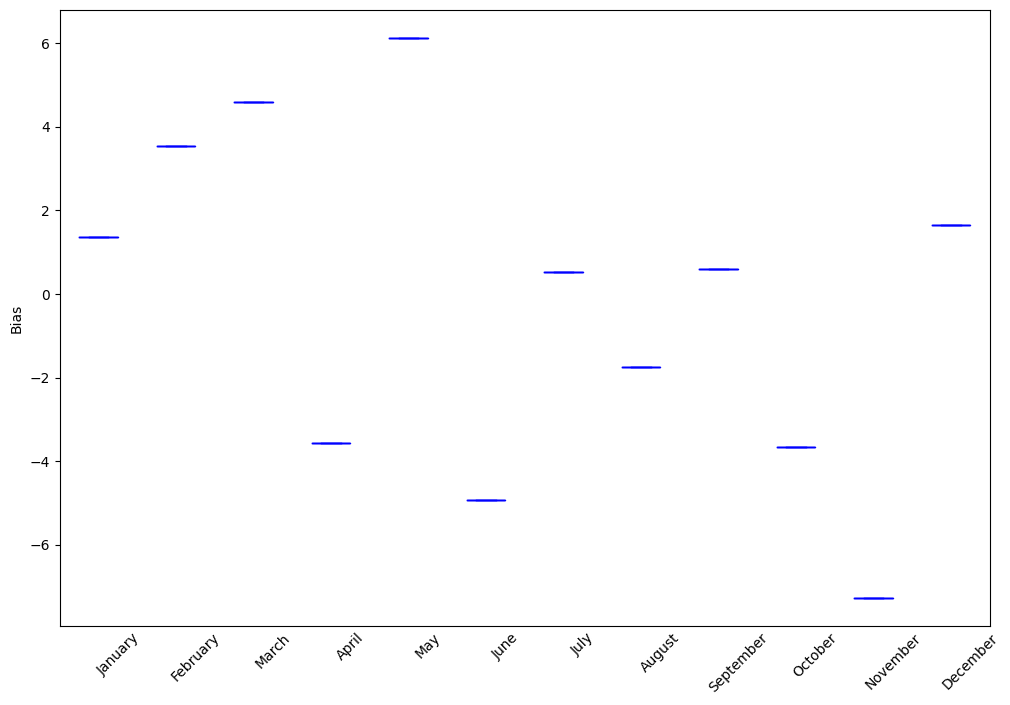
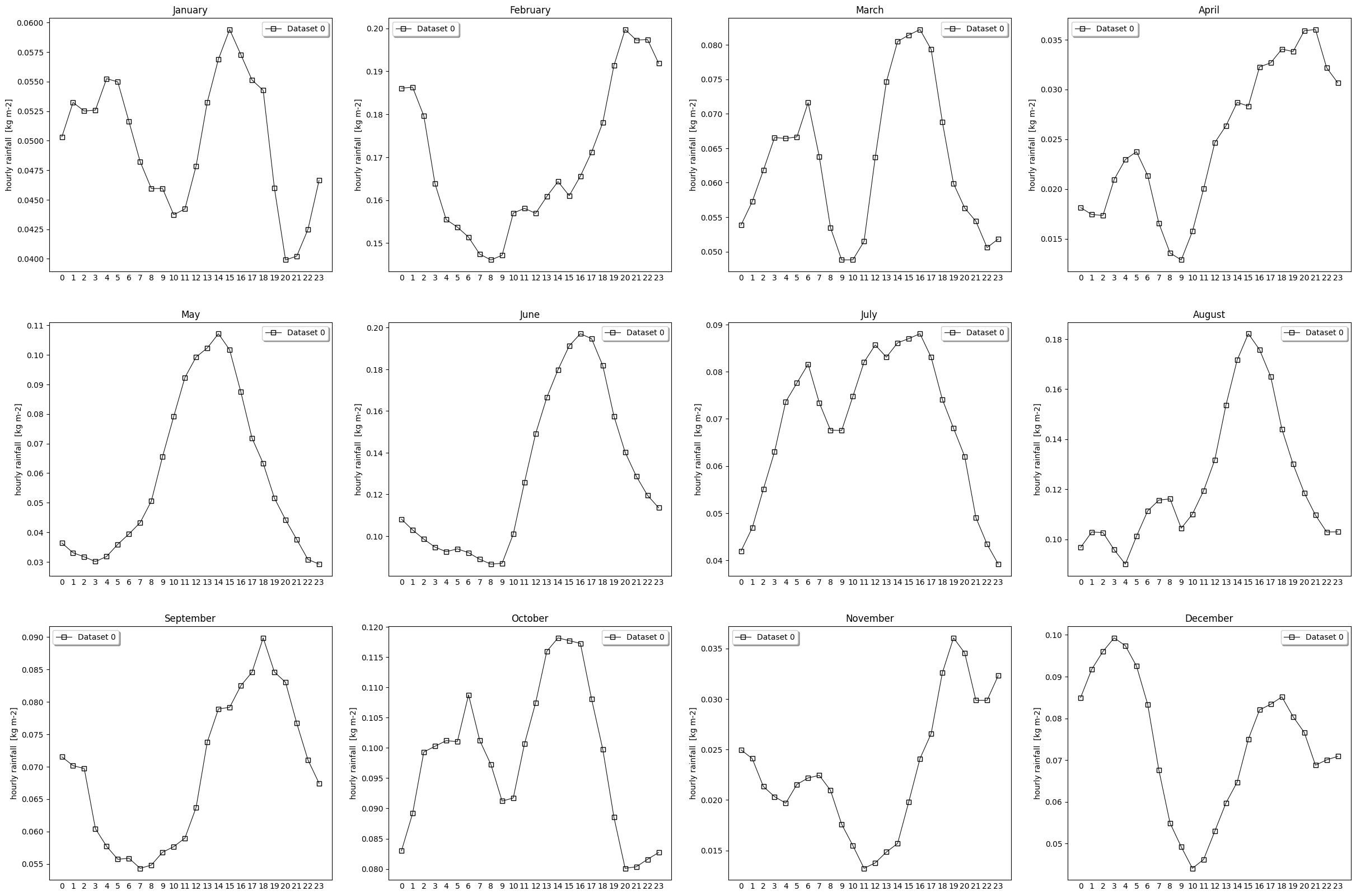
- pyku.analyse.monthly_diurnal_cycle(dat1, dat2=None, var=None, ax=None, **kwargs)[source]#
Diurnal cycle mean for each month
- Parameters:
ds_mod (
xarray.Dataset) – The input dataset.ds_obs (
xarray.Dataset) – Optional. The second dataset.var (str) – variable name
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('low-res-hourly-tas-data') ...: ds = ds.compute() ...: ds.ana.monthly_diurnal_cycle(var='RR') ...: CPU times: user 8.63 s, sys: 6.2 s, total: 14.8 s Wall time: 13 s
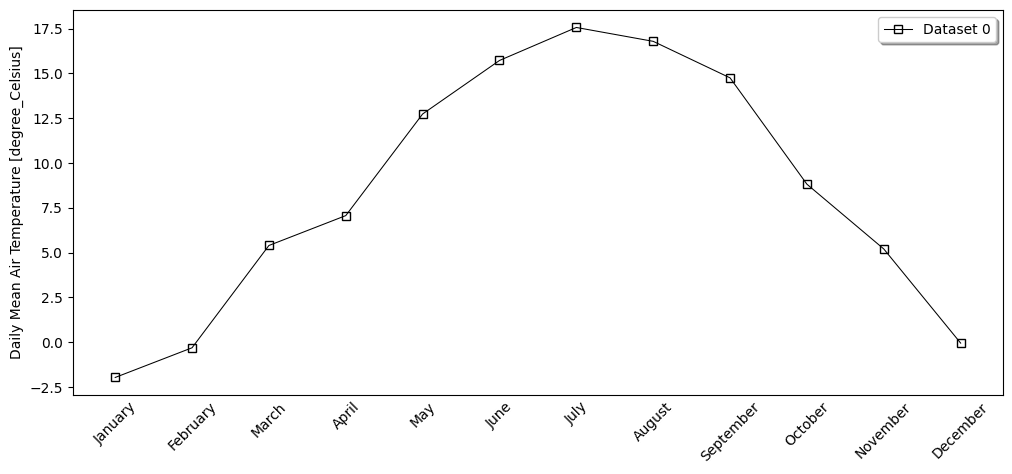
- pyku.analyse.monthly_mean(*dats, var=None, **kwargs)[source]#
Monthly mean
- Parameters:
dats (Union[
xarray.Dataset, List[xarray.Dataset]]) – The input dataset(s)var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas') ...: ds.ana.monthly_mean(var='tas') ...: CPU times: user 680 ms, sys: 194 ms, total: 874 ms Wall time: 877 ms
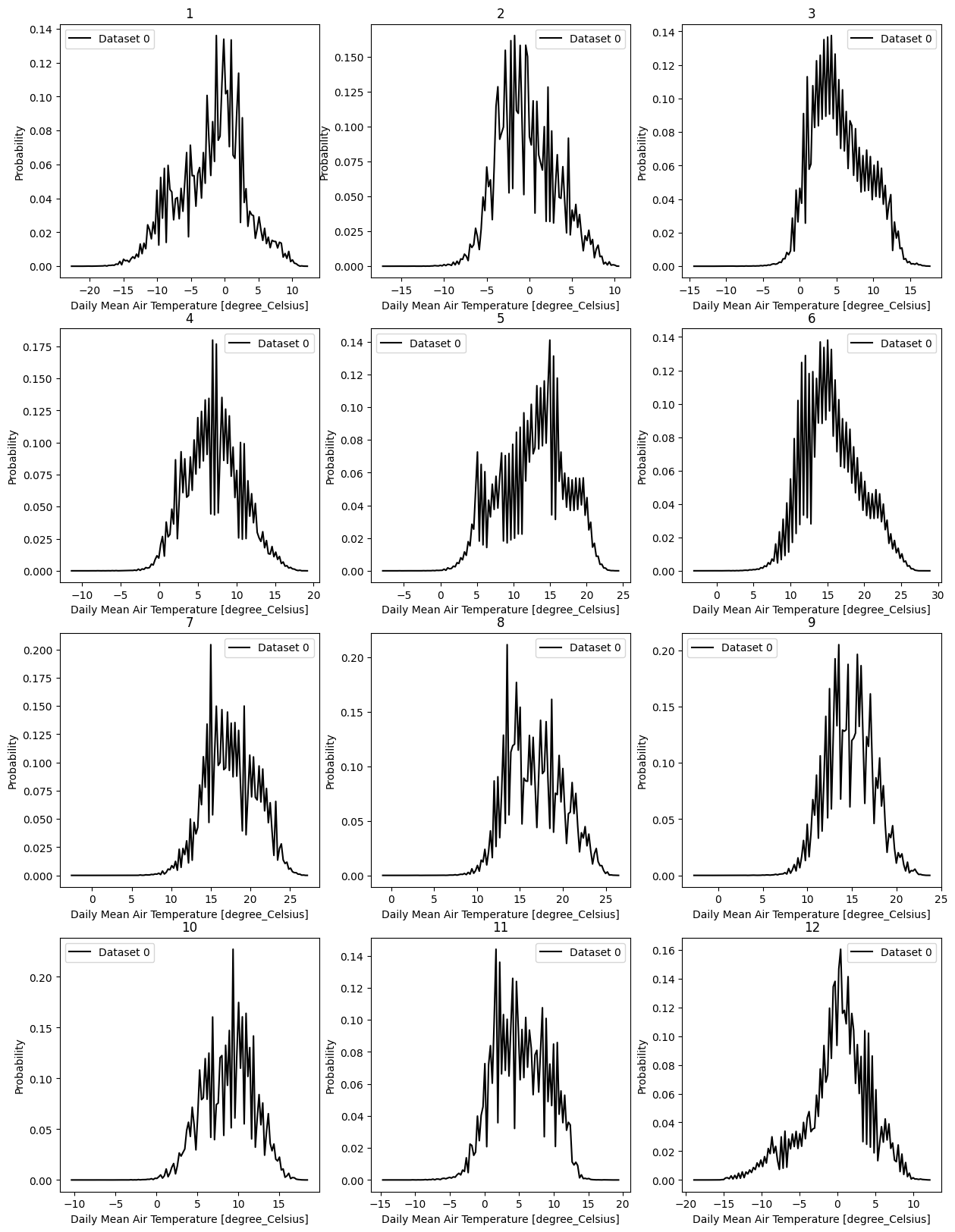
- pyku.analyse.monthly_pdf(*dats, var=None, **kwargs)[source]#
Probability distribution function for each season
- Parameters:
*dats (
xarray.Dataset, List[xarray.Dataset]) – The input dataset(s)var (str) – The variable name.
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – (Optional). The coordinate reference system of the plot.**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas').compute() ...: ds.ana.monthly_pdf(var='tas') ...: CPU times: user 2.84 s, sys: 2.54 s, total: 5.38 s Wall time: 5.21 s
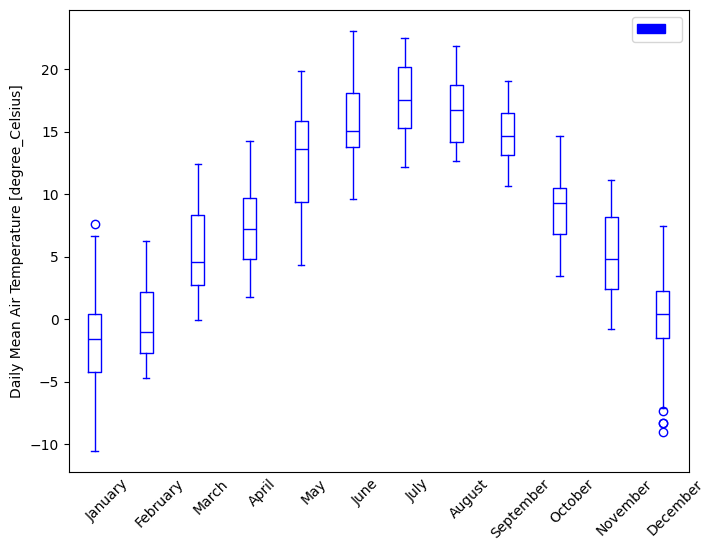
- pyku.analyse.monthly_variability(dat1, dat2=None, var=None, ax=None, **kwargs)[source]#
Monthly variability
- Parameters:
dat1 (
xarray.Dataset) – The first dataset.dat2 (
xarray.Dataset) – Optional. The second dataset.var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas').compute() ...: ds.ana.monthly_variability(var='tas', size_inches=(8, 6)) ...:
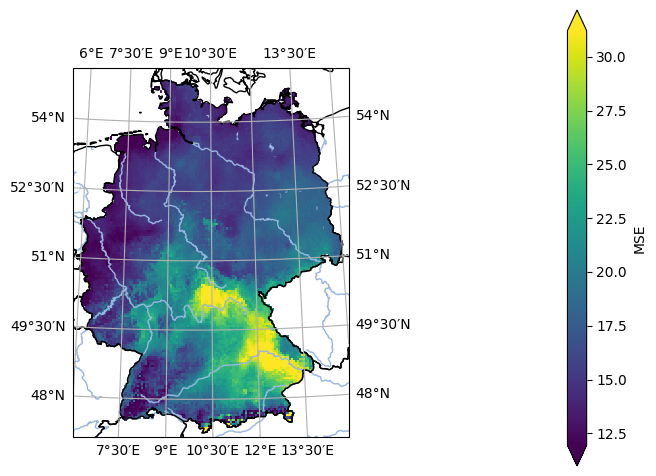
- pyku.analyse.mse_map(*dats, ref, var=None, ds_ref=None, crs=None, **kwargs)[source]#
Map of the Mean Squared Error (MSE)
- Parameters:
dats ([
xarray.Dataset, List[xarray.Dataset]) – The input dataset(s).ref (
xarray.Dataset) – The reference dataset.var (str) – The variable name.
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – Optional. The coordinate reference system of the plot.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Example
In [1]: %%time ...: import pyku ...: ...: # Open reference dataset and preprocess as needed ...: # ----------------------------------------------- ...: ...: ref = ( ...: pyku.resources.get_test_data('hyras') ...: .pyku.to_cmor_units() ...: .pyku.set_time_labels_from_time_bounds(how='lower') ...: .pyku.project('HYR-GER-LAEA-5') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Open model dataset and preprocess as needed ...: # ------------------------------------------- ...: ...: ds = ( ...: pyku.resources.get_test_data('model_data') ...: .pyku.project('HYR-GER-LAEA-5') ...: .pyku.set_time_labels_from_time_bounds(how='lower') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.mse_map(ref=ref, var='tas', crs='HYR-GER-LAEA-5') ...: CPU times: user 1.16 s, sys: 354 ms, total: 1.51 s Wall time: 1.11 s
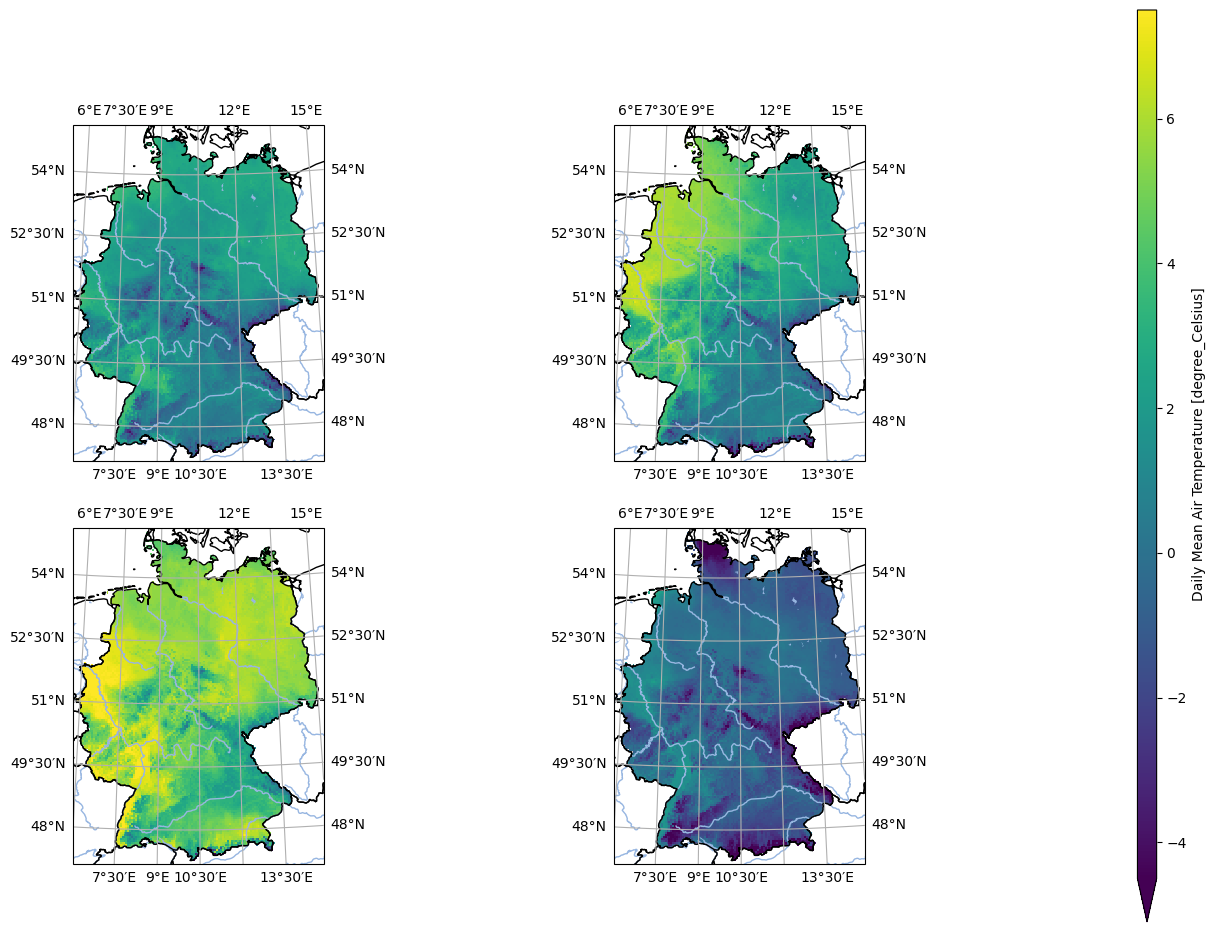
- pyku.analyse.n_maps(*dats, var=None, crs=None, colorbar=True, **kwargs)[source]#
Plot n maps side by side
- Parameters:
dats (
xarray.Dataset, or List[xarray.Dataset]) – The input dataset.var (str) – The variable name
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – (Optional) Coordinate reference system of the plot. If a string is passed, a pre-configured projection definition is taken from the configuration files (e.g. ‘EUR-44’, ‘HYR-LAEA-5’).colorbar (bool) – Optional. Defaults to
True. If set toFalse, the colorbar is not shown and the colormap of all plots is set independently.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed. Further options to control the layout:wspace(value for plt.subplots_adjust),
Example
In [1]: %%time ...: import pyku ...: ...: ds = ( ...: pyku.resources.get_test_data('small_tas_dataset') ...: .compute() ...: ) ...: ...: pyku.analyse.n_maps( ...: ds.isel(time=0), ...: ds.isel(time=1), ...: ds.isel(time=2), ...: ds.isel(time=3), ...: nrows=2, ...: ncols=2, ...: var='tas', ...: crs='HYR-GER-LAEA-5' ...: ) ...: CPU times: user 328 ms, sys: 257 ms, total: 585 ms Wall time: 380 ms
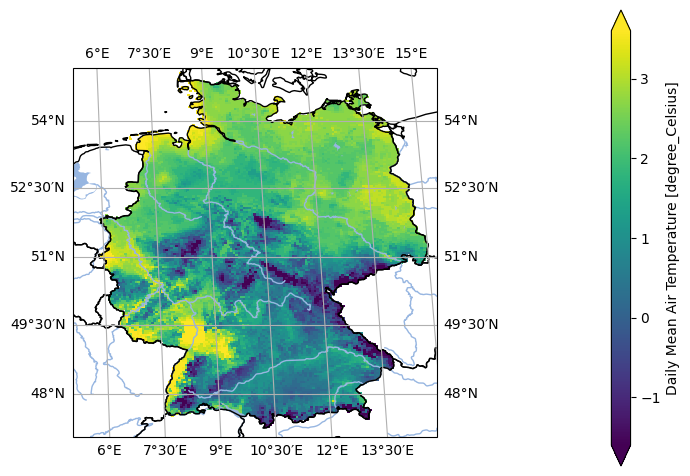
- pyku.analyse.one_map(dat, var=None, crs=None, **kwargs)[source]#
Plot one map
- Parameters:
dat (
xarray.Dataset) – The input dataset.var (str) – The name of the variable.
crs (Union[
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str]) – Coordinate reference system. A pyku pre-defined projection identifier can be passed with a string.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes(),matplotlib.pyplot.figure(), orcartopy.mpl.geoaxes.GeoAxes.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Example
In [1]: %%time ...: import pyku ...: ds = ( ...: pyku.resources.get_test_data('small_tas_dataset') ...: .isel(time=0).compute() ...: ) ...: ds.ana.one_map(var='tas', crs='seamless_world') ...: CPU times: user 219 ms, sys: 199 ms, total: 419 ms Wall time: 271 ms
- pyku.analyse.p98_map(*dats, var, crs=None, **kwargs)[source]#
Map of the 98th percentile
- Parameters:
*dats ([
xarray.Dataset, List[:class:`xarray.Dataset]]) – The input dataset(s)var (str) – The variable name.
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – Optional. The coordinate reference system of the plot.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
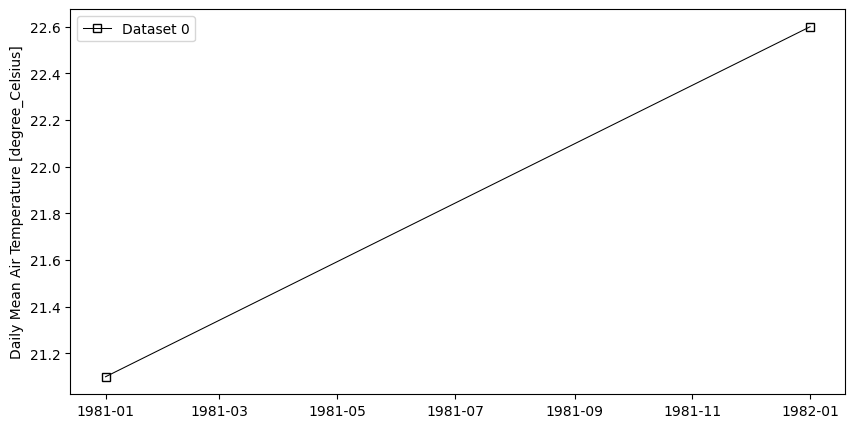
- pyku.analyse.p98_vs_time(*dats, var=None, time_resolution='1YS', ax=None, **kwargs)[source]#
98th percentile vs time
- Parameters:
*dats ([
xarray.Dataset, List[xarray.Dataset]]) – The input dataset(s).time_resolution (str) – The time resolution, e.g. ‘1MS’ or ‘6D’.
var (str) – The variable name.
( (ax) – class:matplotlib.pyplot.axes): Optional. Matplotlib pyplot axis.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas') ...: ds.ana.p98_vs_time(var='tas') ...: CPU times: user 485 ms, sys: 157 ms, total: 642 ms Wall time: 642 ms
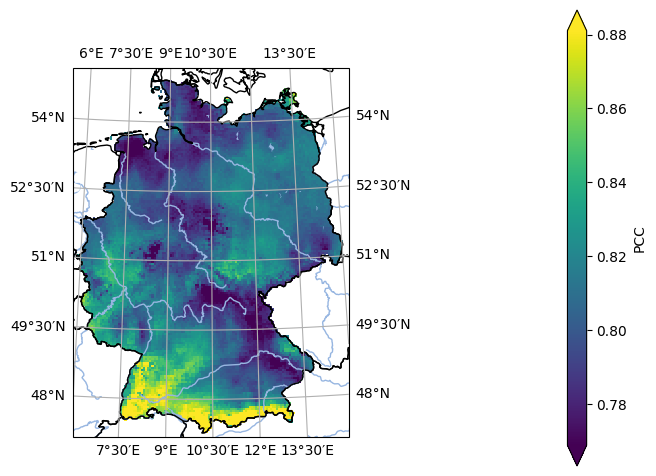
- pyku.analyse.pcc_map(*dats, ref, var=None, crs=None, **kwargs)[source]#
Map of the Pearson Correlation Coefficient (PCC)
- Parameters:
dats ([
xarray.Dataset, List[xarray.Datasets]]) – The input dataset(s)ref (
xarray.Dataset) – The reference dataset.var (str) – Variable name
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – Optional. The coordinate reference system of the plot.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Example
In [1]: %%time ...: import pyku ...: ...: # Open reference dataset and preprocess as needed ...: # ----------------------------------------------- ...: ...: ref = ( ...: pyku.resources.get_test_data('hyras') ...: .pyku.to_cmor_units() ...: .pyku.set_time_labels_from_time_bounds(how='lower') ...: .pyku.project('HYR-GER-LAEA-5') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Open model dataset and preprocess as needed ...: # ------------------------------------------- ...: ...: ds = ( ...: pyku.resources.get_test_data('model_data') ...: .pyku.project('HYR-GER-LAEA-5') ...: .pyku.set_time_labels_from_time_bounds(how='lower') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.pcc_map(ref=ref, var='tas', crs='HYR-GER-LAEA-5') ...: CPU times: user 1.15 s, sys: 380 ms, total: 1.53 s Wall time: 1.13 s
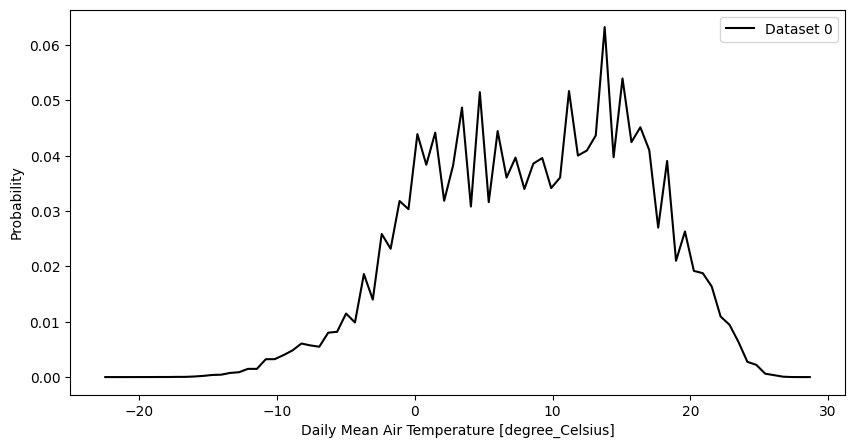
- pyku.analyse.pdf(*dats, var=None, ax=None, **kwargs)[source]#
Probability distribution function
- Parameters:
dats ([
xarray.Dataset, List[xarray.DataArray]]) – The input dataset(s)var (str) – The variable names.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figureormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.range (tuple) – data Range, e.g.
range=(0, 10).nbins (int) – Number of bins.
nsamples (int) – Number of points sampled from the data..
Example
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas').compute() ...: ds.ana.pdf(var='tas', nbins=80) ...: CPU times: user 338 ms, sys: 111 ms, total: 449 ms Wall time: 445 ms
- pyku.analyse.pdf2D(ds, var=None, varx=None, vary=None, sample_size=100, ax=None, **kwargs)[source]#
2D distribution
- Parameters:
ds (
xarray.Dataset) – The input dataset.varx (str) – The variable on the x axis.
vary (str) – The variable on the y axis.
sample_size (int) – The sample size.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
- pyku.analyse.percentile_map(ds_mod, ds_obs, var=None, percentile=0.5, crs=None, **kwargs)[source]#
Map of the nth percentile
Todo
Check if function is actually working
Create a documentation
I think the CRS should also work with
pyresample.geometry.AreaDefinition.
- Parameters:
ds_mod (
xarray.Dataset) – The first dataset.ds_obs (
xarray.Dataset) – The second dataset.var (str) – Variable name
percentile (float) – Percentile between 0 and 1
crs (
cartopy.crs.CRS) – Coordinate reference system
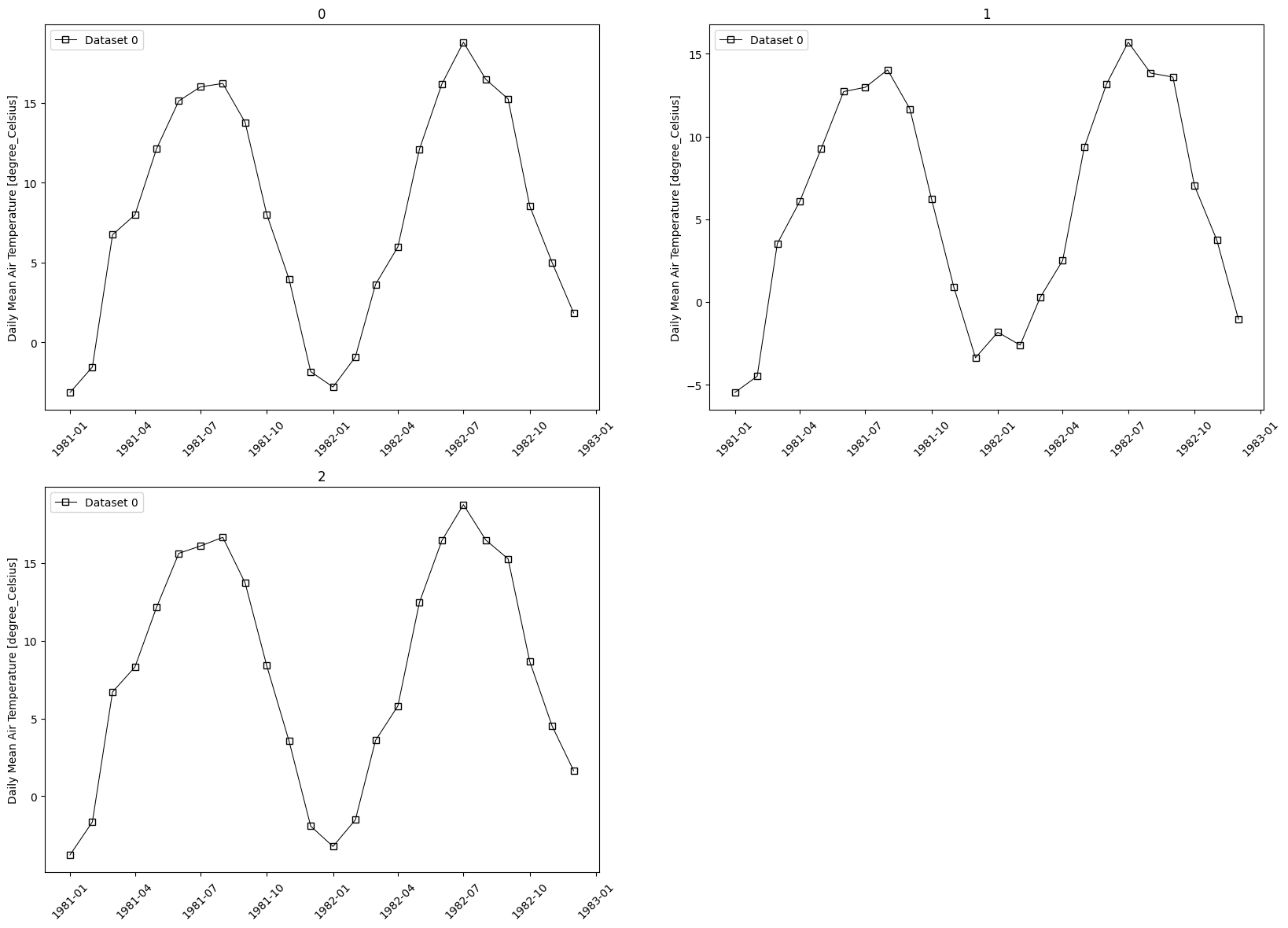
- pyku.analyse.regional_mean_vs_time(ds_mod, ds_obs=None, gdf=None, time_resolution='1MS', var=None, **kwargs)[source]#
Time series per region
- Parameters:
ds_mod (
xarray.Dataset) – The first input dataset.ds_obs (
xarray.Dataset) – The second input dataset.gdf (
geopandas.GeoDataFrame) – The polygon data.var (str) – variable name
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Examples
In [1]: %%time ...: import xarray as xr ...: import pyku ...: import pyku.analyse as analyse ...: ...: # Open and preprocess test data as needed ...: # --------------------------------------- ...: ...: ds = pyku.resources.get_test_data('hyras') ...: ...: # Get a subset ot the natrual areas or germany ...: # -------------------------------------------- ...: ...: regions = ( ...: pyku.resources ...: .get_geodataframe('natural_areas_of_germany') ...: .head(3) ...: ) ...: ...: ds.ana.regional_mean_vs_time( ...: gdf=regions, ...: time_resolution='1MS', ...: var='tas' ...: ) ...: CPU times: user 11.6 s, sys: 2.1 s, total: 13.7 s Wall time: 13.6 s
- pyku.analyse.regional_monthly_bias(ds_mod, ds_obs, gdf, area_def, var=None, **kwargs)[source]#
Monthly variability
Todo
Check if this function is working
Get the ipython code right
Check documentation is right. I think area_def can be passed as cartopy crs?
- Parameters:
ds_mod (
xarray.Dataset) – The first input dataset.ds_obs (
xarray.Dataset) – The second input dataset.gdf (
geopandas.GeoDataFrame) – The polygon data.var (str) – variable name
area_def (
pyresample.geometry.AreaDefinition) – Optional. The plot coordinate reference system.**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Examples
In [1]: %%time ...: import pyku ...: ...: # Open reference data and preprocess as needed ...: # -------------------------------------------- ...: ...: ref = ( ...: pyku.resources.get_test_data('hyras') ...: .pyku.to_cmor_units() ...: .pyku.set_time_labels_from_time_bounds(how='lower') ...: .pyku.project('HYR-LAEA-50') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Open model data and preprocess as needed ...: # ---------------------------------------- ...: ...: ds = ( ...: pyku.resources.get_test_data('model_data') ...: .pyku.project('HYR-LAEA-50') ...: .pyku.set_time_labels_from_time_bounds(how='lower') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: gdf = ( ...: pyku.resources ...: .get_geodataframe('natural_areas_of_germany') ...: .loc[0:4] ...: ) ...: ...: print('Polygons: ', gdf) ...: ...: # Calculate and plot ...: # ------------------ ...: ...: # pyku.analyse.regional_monthly_bias( ...: # ds, ref, gdf, area_def='HYR-LAEA-50', var='tas' ...: # ) ...: Polygons: Shape_Leng ... geometry 0 1.676004e+06 ... POLYGON ((3613725.5 5598209, 3613437 5597195, ... 1 8.566480e+05 ... MULTIPOLYGON (((3614073 5273052, 3614476 52725... 2 1.096050e+06 ... POLYGON ((3734247 5437391, 3734470 5437339, 37... 3 9.577557e+05 ... POLYGON ((3288568.5 5633985.5, 3288857.25 5633... 4 1.457899e+06 ... MULTIPOLYGON (((3794440.445 6030118.175, 37943... [5 rows x 12 columns] CPU times: user 870 ms, sys: 183 ms, total: 1.05 s Wall time: 819 ms
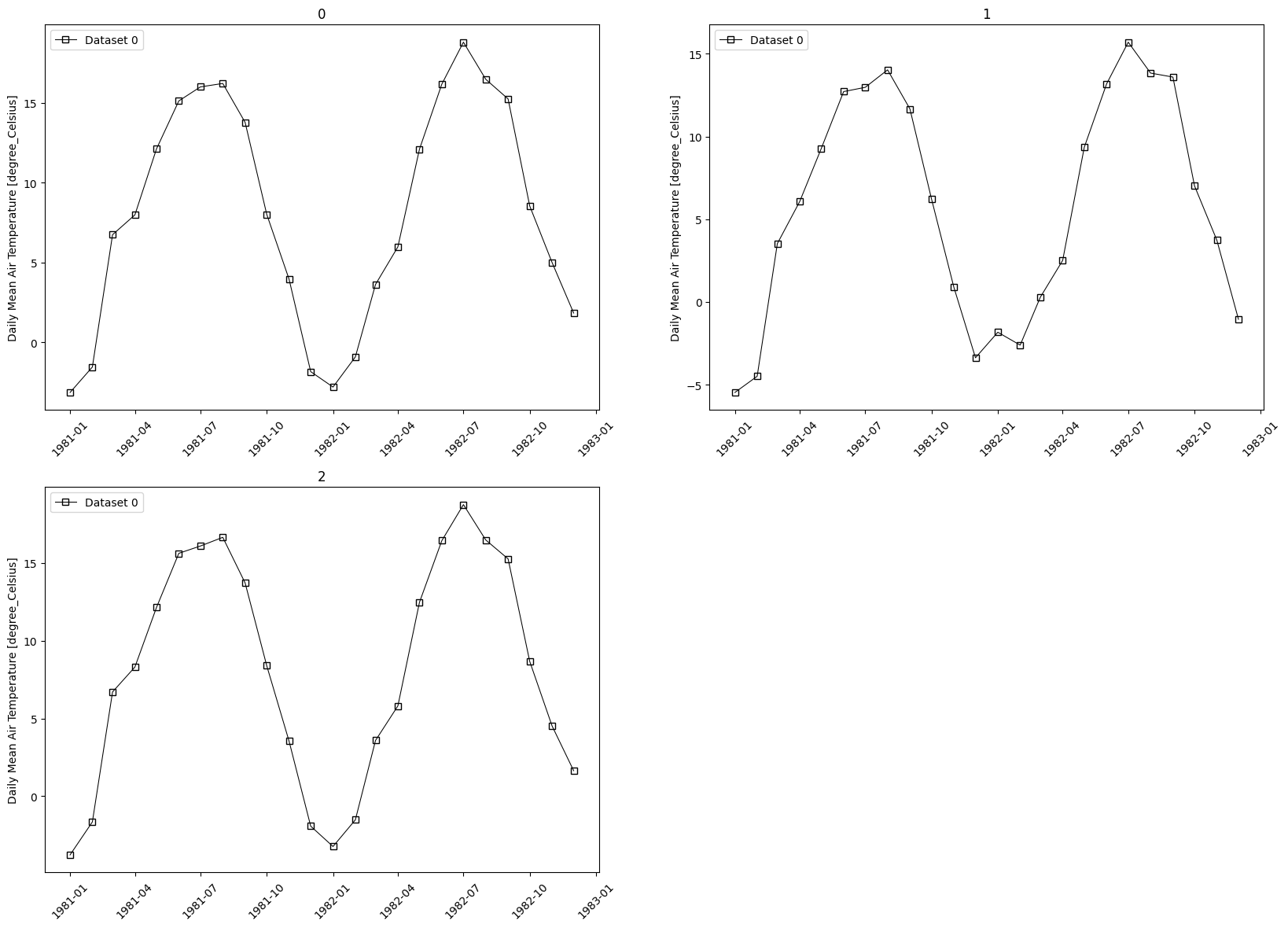
- pyku.analyse.regional_monthly_variability(ds1, ds2, gdf=None, var=None, **kwargs)[source]#
Monthly variability
- Parameters:
ds1 (
xarray.Dataset) – The first dataset.ds2 (
xarray.Dataset) – The second dataset.gdf (
geopandas.GeoDataFrame) – The polygon data.var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Examples
In [1]: import xarray as xr ...: import pyku ...: import pyku.analyse as analyse ...: ...: # Open and preprocess test data as needed ...: # --------------------------------------- ...: ...: ds1 = ( ...: pyku.resources.get_test_data('hyras') ...: .pyku.to_cmor_varnames() ...: .pyku.to_cmor_units() ...: ) ...: ...: ds2 = pyku.resources.get_test_data('cordex_data') ...: ...: # Get a subset ot the natrual areas or germany ...: # -------------------------------------------- ...: ...: regions = ( ...: pyku.resources ...: .get_geodataframe('natural_areas_of_germany') ...: .head(3) ...: ) ...: ...: # Calculate and plot ...: # ------------------ ...: ...: analyse.regional_monthly_variability( ...: ds1, ...: ds2, ...: gdf=regions, ...: var='tas' ...: ) ...:
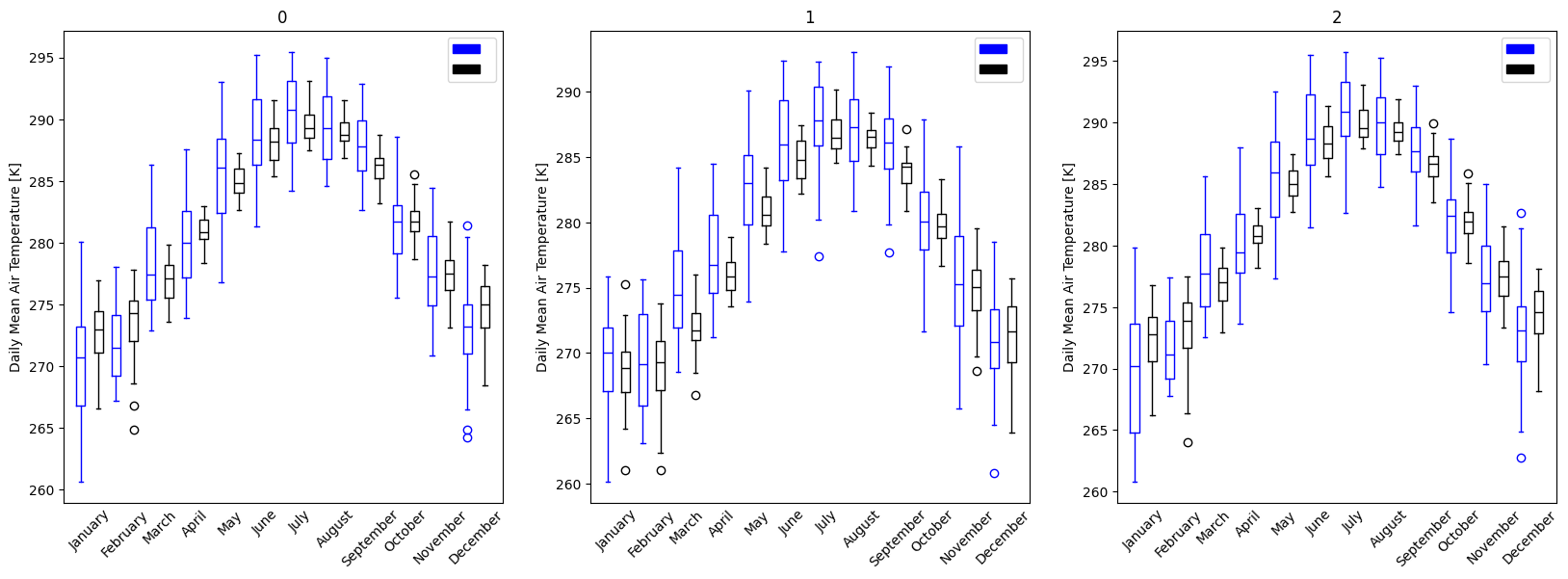
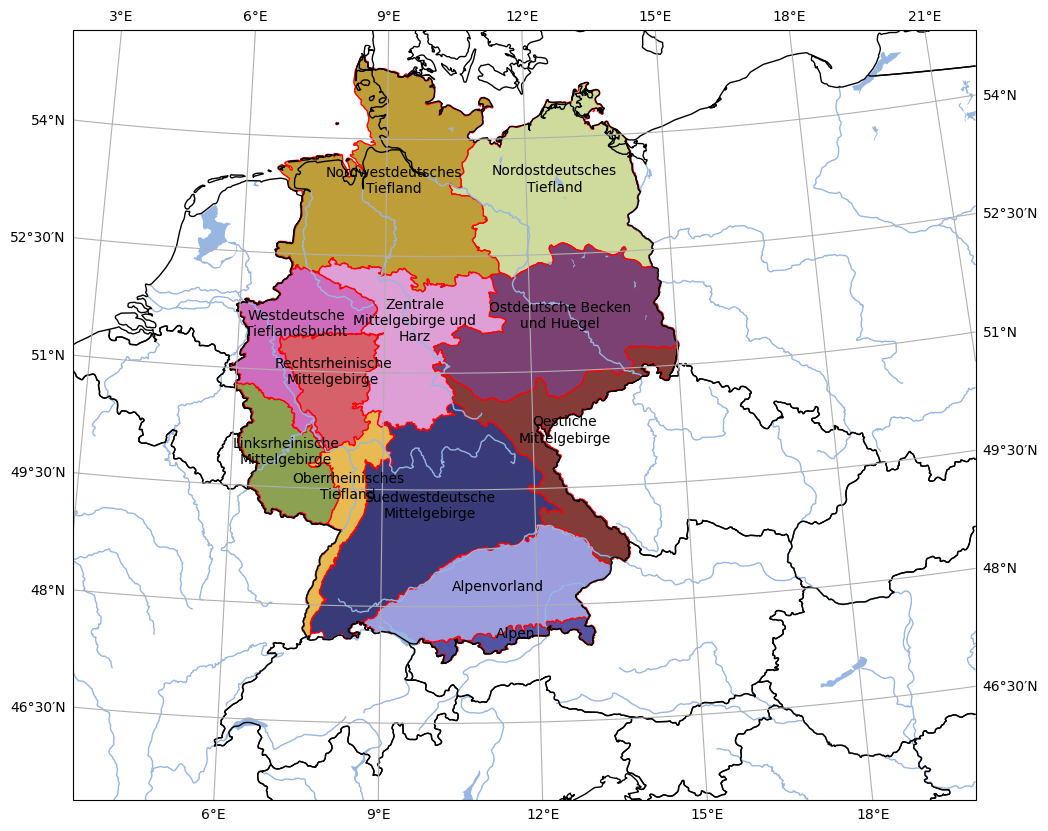
- pyku.analyse.regions(gdf, area_def, **kwargs)[source]#
Plot regions
- Parameters:
gdf (geopandas.GeoDataFrame) – Polygon data
area_def ([pyresample.AreaDefinition, string]) – Area definition
**kwargs – Any argument that can be passed to
matplotlib.ArtistIn particularsize_inchescan be passed.
Example
In [1]: import pyku ...: gdf = pyku.resources.get_geodataframe( ...: 'natural_areas_of_germany' ...: ) ...: pyku.analyse.regions(gdf, area_def='HYR-LAEA-5') ...:
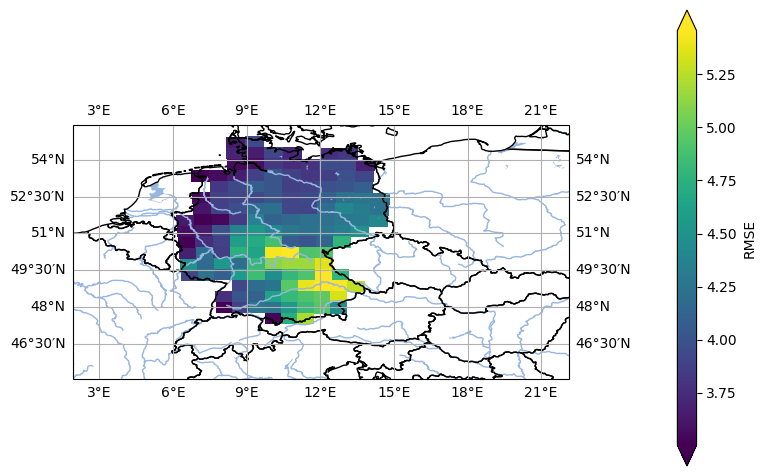
- pyku.analyse.rmse_map(*dats, ref, var=None, crs=None, ds_ref=None, **kwargs)[source]#
Map of the Root Mean Squared Error (RMSE)
- Parameters:
dats ([
xarray.Dataset, List[xarray.Dataset]]) – The input dataset(s).ref (
xarray.Dataset) – The reference dataset.var (str) – The variable name.
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – Optional. The coordinate reference system of the plot.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Example
In [1]: %%time ...: import pyku ...: ...: # Open reference dataset and preprocess as needed ...: # ----------------------------------------------- ...: ...: ref = pyku.resources.get_test_data('hyras')\ ...: .pyku.to_cmor_units()\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .sel(time='1981')\ ...: .load() ...: ...: # Open model dataset and preprocess as needed ...: # ------------------------------------------- ...: ...: ds = pyku.resources.get_test_data('model_data')\ ...: .pyku.project('HYR-LAEA-50')\ ...: .pyku.set_time_labels_from_time_bounds(how='lower')\ ...: .sel(time='1981')\ ...: .load() ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.rmse_map(ref=ref, var='tas') ...: CPU times: user 1.02 s, sys: 377 ms, total: 1.39 s Wall time: 941 ms
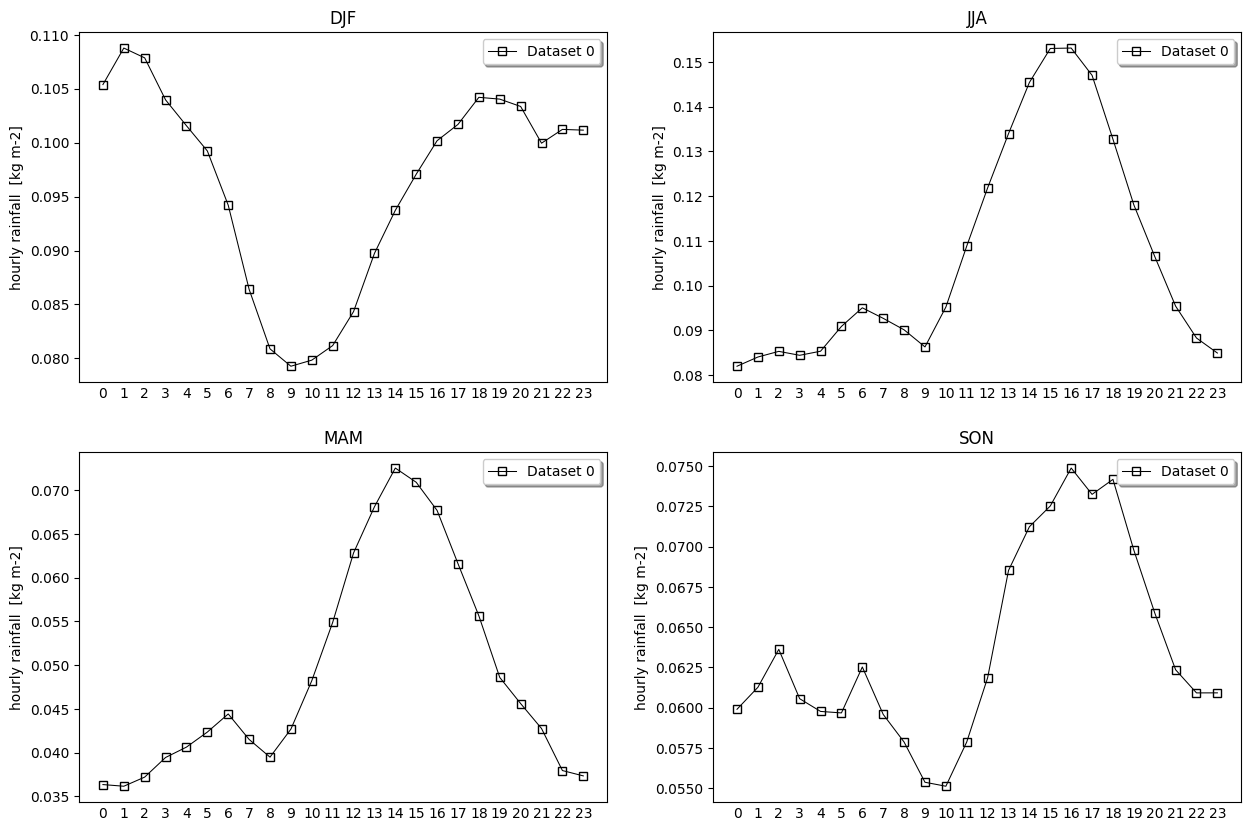
- pyku.analyse.seasonal_diurnal_cycle(dat1, dat2=None, var=None, ax=None, **kwargs)[source]#
Diurnal cycle for each seasons
- Parameters:
ds_mod (
xarray.Dataset) – Model datads_obs (
xarray.Dataset) – Reference datavar (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Examples
In [1]: %%time ...: import pyku ...: ds = pyku.resources.get_test_data('low-res-hourly-tas-data') ...: ds.ana.seasonal_diurnal_cycle(var='RR') ...: CPU times: user 7.78 s, sys: 2.92 s, total: 10.7 s Wall time: 8.76 s
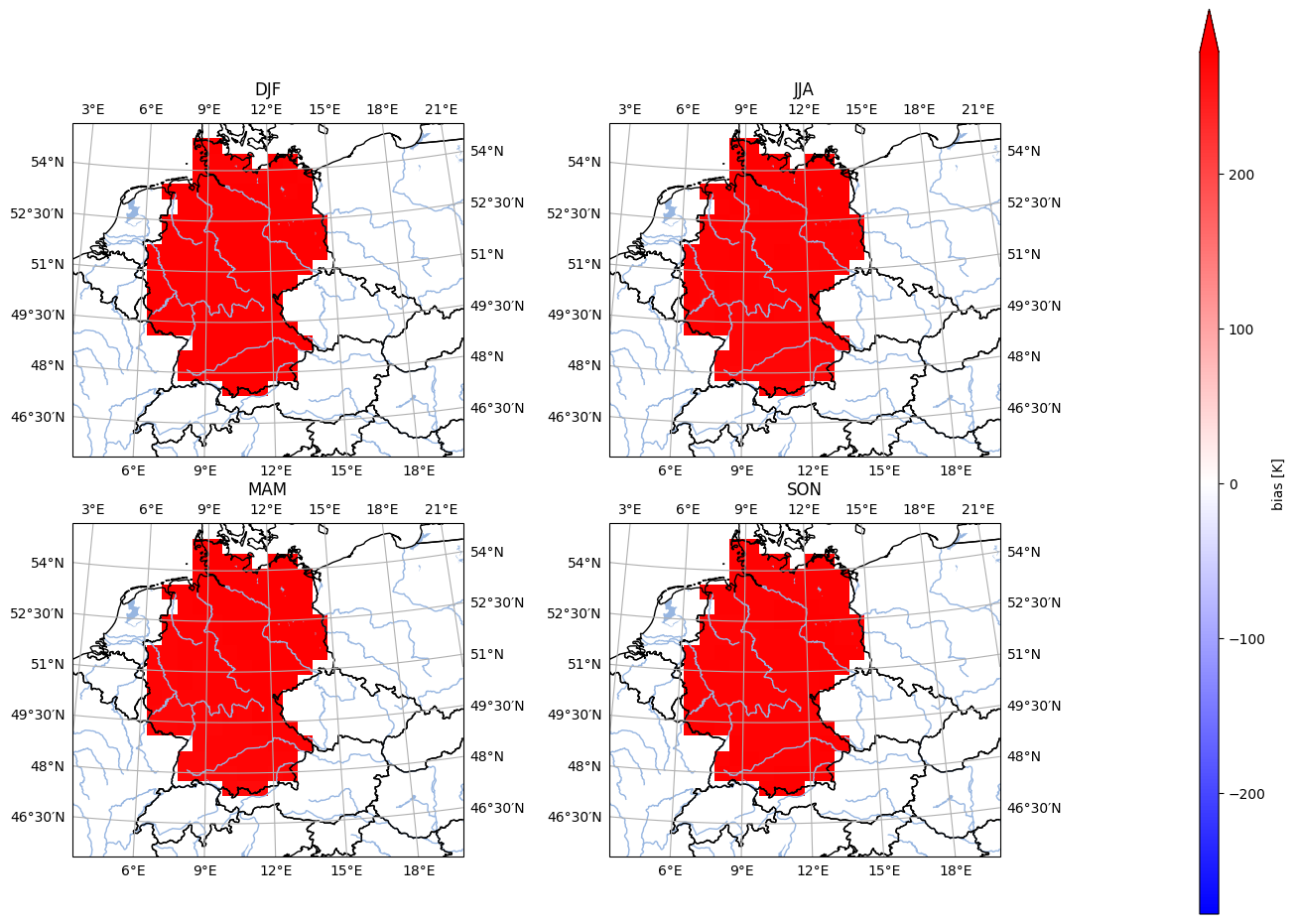
- pyku.analyse.seasonal_mean_bias_map(dats, ref, var=None, crs=None, **kwargs)[source]#
Map of the mean bias
- Parameters:
dats (
xarray.Dataset) – The input model datasetref (
xarray.Dataset) – The Reference/Observation datavar (str) – The variable name
crs (
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str) – Optional. The plot coordinate reference system.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.gridliner.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Notes
If an argument is passed which is not recognized, no error message is thrown and the argument is ignored.
Example
In [1]: import pyku ...: ...: # Open reference dataset and preprocess as needed ...: # ----------------------------------------------- ...: ...: ref = ( ...: pyku.resources.get_test_data('hyras') ...: .pyku.project('HYR-LAEA-50') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Open model dataset and preprocess as needed ...: # ------------------------------------------- ...: ...: ds = ( ...: pyku.resources.get_test_data('model_data') ...: .pyku.project('HYR-LAEA-50') ...: .sel(time='1981') ...: .compute() ...: ) ...: ...: # Calculate and plot ...: # ------------------ ...: ...: ds.ana.seasonal_mean_bias_map( ...: ref=ref, ...: var='tas', ...: crs='HYR-LAEA-50' ...: ) ...:
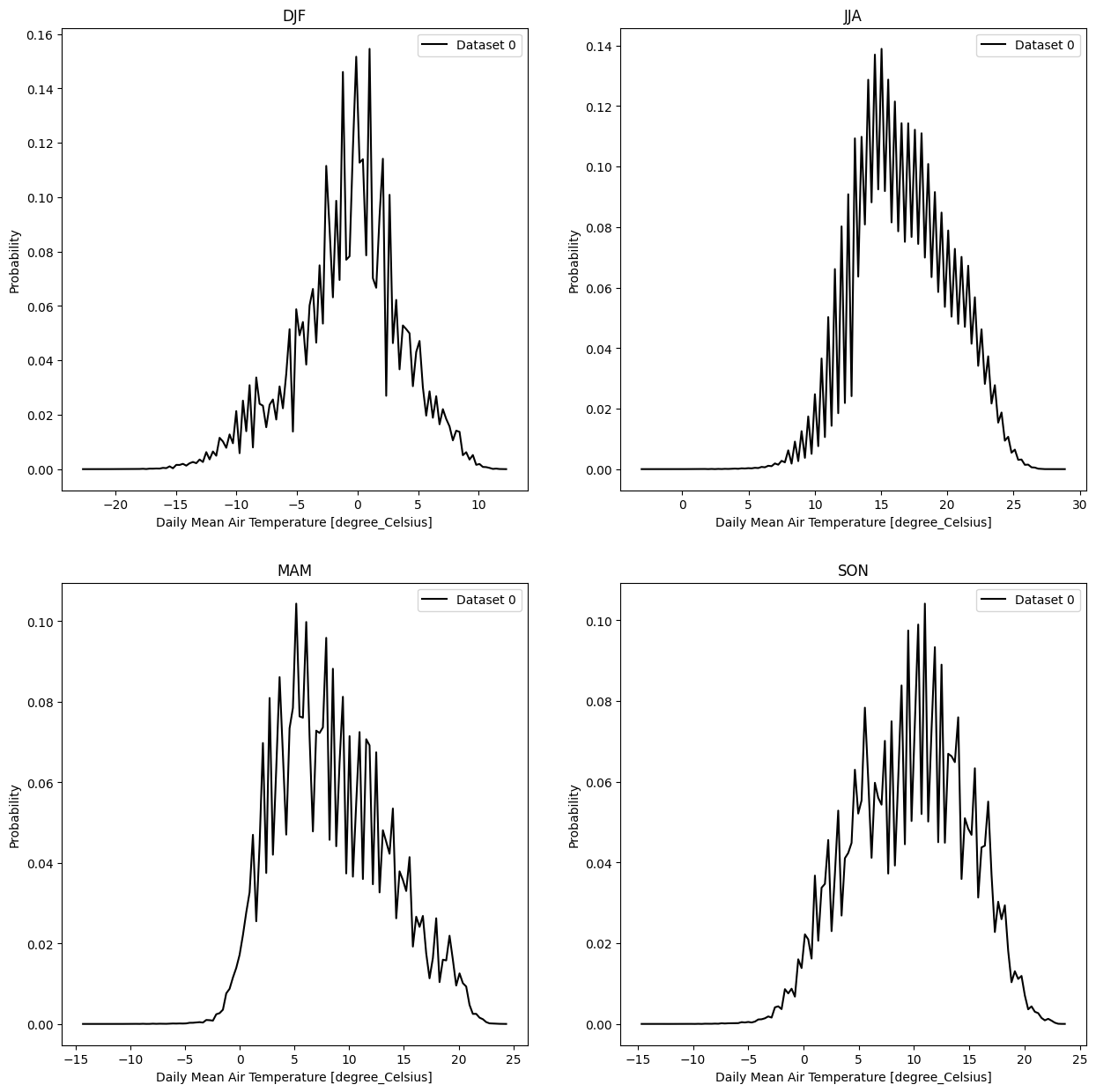
- pyku.analyse.seasonal_pdf(*dats, var=None, **kwargs)[source]#
Probability distribution function for each season
- Parameters:
dats (
xarray.Dataset, List[xarray.Dataset]]) – The input dataset(s).var (str) – The variable name.
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: import pyku ...: ds = pyku.resources.get_test_data('hyras_tas') ...: ds.ana.seasonal_pdf(var='tas') ...:
- pyku.analyse.summary_table(ds_mod, ds_obs, **kwargs)[source]#
Summary in table form
- Parameters:
ds_mod (
xarray.Dataset) – Model dataset.ds_obs (
xarray.Dataset) – Reference dataset.
- Returns:
reStructuredText table
- Return type:
str
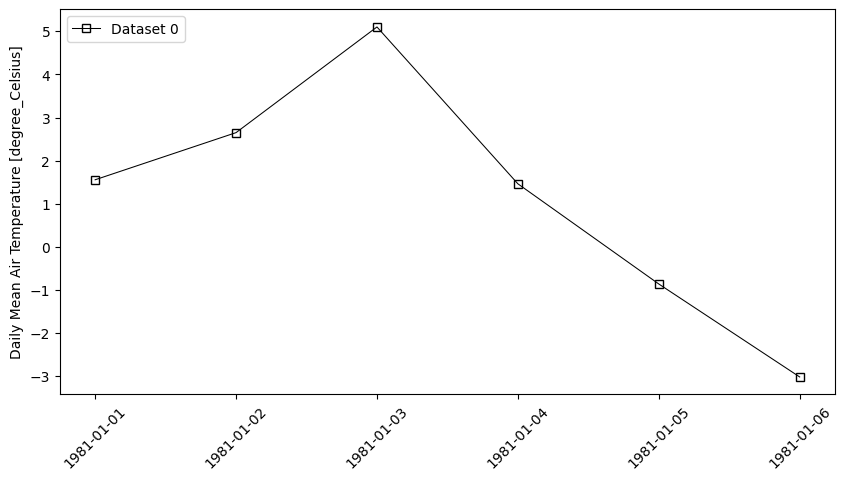
- pyku.analyse.time_serie(*dats, var=None, ax=None, **kwargs)[source]#
Time serie. The data are averaged over y/x and plotting against time
- Parameters:
*dats (
xarray.Dataset, List[xarray.Dataset]) – The input dataset(s).var (str) – The variable name.
ax (
matplotlib.pyplot.axes) – Optional. Matplotlib pyplot axis.**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
Example
In [1]: pyku ...: ds = pyku.resources.get_test_data('hyras_tas')\ ...: .isel(time=[0,1,2,3,4,5]) ...: ds.ana.time_serie(var='tas') ...:
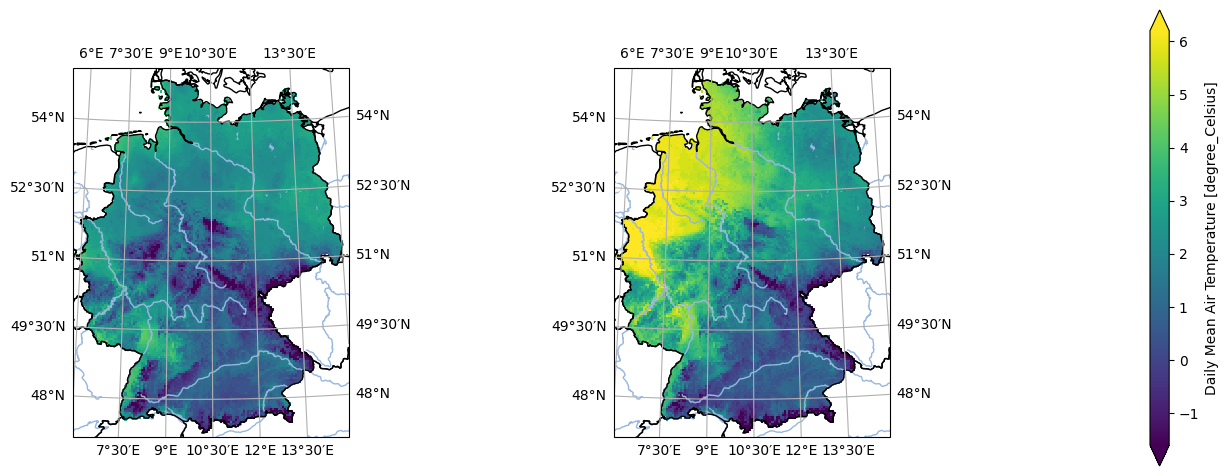
- pyku.analyse.two_maps(dat1, dat2, var=None, crs=None, **kwargs)[source]#
Plot two maps side by side
- Parameters:
dat1 (
xarray.Dataset) – The first dataset.dat2 (
xarray.Dataset) – The second dataset.var (str) – The name of the variable.
crs (Union[
cartopy.crs.CRS,pyresample.geometry.AreaDefinition, str]) – Coordonate reference system. A pyku pre-defined projection identifier can be passed with a string.**kwargs – Any argument that can be passed to
matplotlib.pyplot.pcolormesh(),matplotlib.pyplot.axes()matplotlib.pyplot.figure(), orcartopy.mpl.geoaxes.GeoAxes.gridlines(). For example,cmap,vmin,vmax, orsize_inchescan be passed.
Notes
If an argument is passed which is not recognized, no error message is thrown and the argument is ignored.
Example
In [1]: %%time ...: import pyku ...: ...: ds = pyku.resources.get_test_data('small_tas_dataset') ...: ...: pyku.analyse.two_maps( ...: ds.isel(time=0), ...: ds.isel(time=1), ...: rows=1, ...: cols=2, ...: var='tas', ...: crs='HYR-GER-LAEA-5', ...: ) ...: CPU times: user 253 ms, sys: 203 ms, total: 457 ms Wall time: 284 ms
- pyku.analyse.var_vs_year(dat_mod, dat_obs, var=None, **kwargs)[source]#
Variability vs year
Todo
Function looks very outdated and needs to be reworked
- Parameters:
ds_mod (
xarray.Dataset) – The first dataset.ds_obs (
xarray.Dataset) – The second dataset.var (str) – Variable name
**kwargs – Any argument that can be passed to
matplotlib.pyplot.figure()ormatplotlib.pyplot.axes(). In particularylim=(0,15),size_inches=(20, 40)), oryscale='log'can be passed.
